Columbia Data Science course, week 10: Observational studies, confounders, epidemiology
This week our guest lecturer in the Columbia Data Science class was David Madigan, Professor and Chair of Statistics at Columbia. He received a bachelors degree in Mathematical Sciences and a Ph.D. in Statistics, both from Trinity College Dublin. He has previously worked for AT&T Inc., Soliloquy Inc., the University of Washington, Rutgers University, and SkillSoft, Inc. He has over 100 publications in such areas as Bayesian statistics, text mining, Monte Carlo methods, pharmacovigilance and probabilistic graphical models.
So Madigan is an esteemed guest, but I like to call him an “apocalyptic leprechaun”, for reasons which you will know by the end of this post. He’s okay with that nickname, I asked his permission.
Madigan came to talk to us about observation studies, of central importance in data science. He started us out with this:
Thought Experiment
We now have detailed, longitudinal medical data on tens of millions of patients. What can we do with it?
To be more precise, we have tons of phenomenological data: this is individual, patient-level medical record data. The largest of the databases has records on 80 million people: every prescription drug, every condition ever diagnosed, every hospital or doctor’s visit, every lab result, procedures, all timestamped.
But we still do things like we did in the Middle Ages; the vast majority of diagnosis and treatment is done in a doctor’s brain. Can we do better? Can you harness these data to do a better job delivering medical care?
Students responded:
1) There was a prize offered on Kaggle, called “Improve Healthcare, Win $3,000,000.” predicting who is going to go to the hospital next year. Doesn’t that give us some idea of what we can do?
Madigan: keep in mind that they’ve coarsened the data for proprietary reasons. Hugely important clinical problem, especially as a healthcare insurer. Can you intervene to avoid hospitalizations?
2) We’ve talked a lot about the ethical uses of data science in this class. It seems to me that there are a lot of sticky ethical issues surrounding this 80 million person medical record dataset.
Madigan: Agreed! What nefarious things could we do with this data? We could gouge sick people with huge premiums, or we could drop sick people from insurance altogether. It’s a question of what, as a society, we want to do.
What is modern academic statistics?
Madigan showed us Drew Conway’s Venn Diagram that we’d seen in week 1:
 Madigan positioned the modern world of the statistician in the green and purple areas.
Madigan positioned the modern world of the statistician in the green and purple areas.
It used to be the case, say 20 years ago, according to Madigan, that academic statistician would either sit in their offices proving theorems with no data in sight (they wouldn’t even know how to run a t-test) or sit around in their offices and dream up a new test, or a new way of dealing with missing data, or something like that, and then they’d look around for a dataset to whack with their new method. In either case, the work of an academic statistician required no domain expertise.
Nowadays things are different. The top stats journals are more deep in terms of application areas, the papers involve deep collaborations with people in social sciences or other applied sciences. Madigan is setting an example tonight by engaging with the medical community.
Madigan went on to make a point about the modern machine learning community, which he is or was part of: it’s a newish academic field, with conferences and journals, etc., but is characterized by what stats was 20 years ago: invent a method, try it on datasets. In terms of domain expertise engagement, it’s a step backwards instead of forwards.
Comments like the above make me love Madigan.
Very few academic statisticians have serious hacking skills, with Mark Hansen being an unusual counterexample. But if all three is what’s required to be called data science, then I’m all for data science, says Madigan.
Madigan’s timeline
Madigan went to college in 1980, specialized on day 1 on math for five years. In final year, he took a bunch of stats courses, and learned a bunch about computers: pascal, OS, compilers, AI, database theory, and rudimentary computing skills. Then came 6 years in industry, working at an insurance company and a software company where he specialized in expert systems.
It was a mainframe environment, and he wrote code to price insurance policies using what would now be described as scripting languages. He also learned about graphics by creating a graphic representation of a water treatment system. He learned about controlling graphics cards on PC’s, but he still didn’t know about data.
Then he got a Ph.D. and went into academia. That’s when machine learning and data mining started, which he fell in love with: he was Program Chair of the KDD conference, among other things, before he got disenchanted. He learned C and java, R and S+. But he still wasn’t really working with data yet.
He claims he was still a typical academic statistician: he had computing skills but no idea how to work with a large scale medical database, 50 different tables of data scattered across different databases with different formats.
In 2000 he worked for AT&T labs. It was an “extreme academic environment”, and he learned perl and did lots of stuff like web scraping. He also learned awk and basic unix skills.
It was life altering and it changed everything: having tools to deal with real data rocks! It could just as well have been python. The point is that if you don’t have the tools you’re handicapped. Armed with these tools he is afraid of nothing in terms of tackling a data problem.
In Madigan’s opinion, statisticians should not be allowed out of school unless they know these tools.
He then went to a internet startup where he and his team built a system to deliver real-time graphics on consumer activity.
Since then he’s been working in big medical data stuff. He’s testified in trials related to medical trials, which was eye-opening for him in terms of explaining what you’ve done: “If you’re gonna explain logistical regression to a jury, it’s a different kind of a challenge than me standing here tonight.” He claims that super simple graphics help.
Carrotsearch
As an aside he suggests we go to this website, called carrotsearch, because there’s a cool demo on it.
What is an observational study?
Madigan defines it for us:
An observational study is an empirical study in which the objective is to elucidate cause-and-effect relationships in which it is not feasible to use controlled experimentation.
In tonight’s context, it will involve patients as they undergo routine medical care. We contrast this with designed experiment, which is pretty rare. In fact, Madigan contends that most data science activity revolves around observational data. Exceptions are A/B tests. Most of the time, the data you have is what you get. You don’t get to replay a day on the market where Romney won the presidency, for example.
Observational studies are done in contexts in which you can’t do experiments, and they are mostly intended to elucidate cause-and-effect. Sometimes you don’t care about cause-and-effect, you just want to build predictive models. Madigan claims there are many core issues in common with the two.
Here are some examples of tests you can’t run as designed studies, for ethical reasons:
- smoking and heart disease (you can’t randomly assign someone to smoke)
- vitamin C and cancer survival
- DES and vaginal cancer
- aspirin and mortality
- cocaine and birthweight
- diet and mortality
Pitfall #1: confounders
There are all kinds of pitfalls with observational studies.
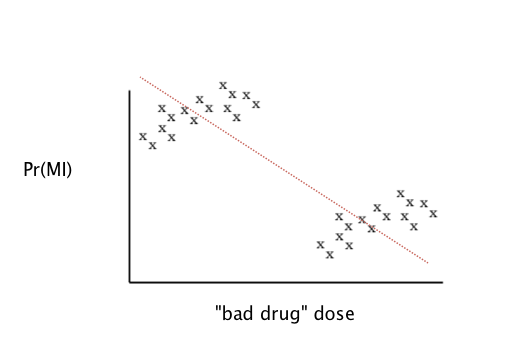
For example, look at this graph, where you’re finding a best fit line to describe whether taking higher doses of the “bad drug” is correlated to higher probability of a heart attack:
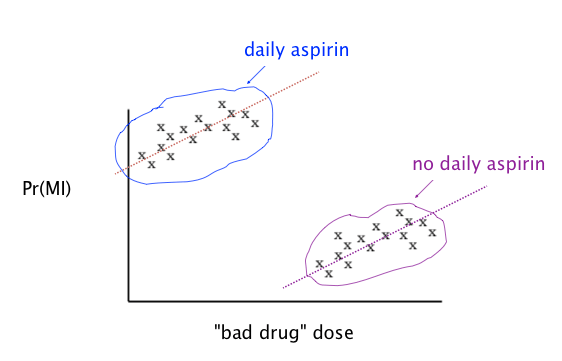
It looks like, from this vantage point, the more drug you take the fewer heart attacks you have. But there are two clusters, and if you know more about those two clusters, you find the opposite conclusion:
Note this picture was rigged it so the issue is obvious. This is an example of a “confounder.” In other words, the aspirin-taking or non-aspirin-taking of the people in the study wasn’t randomly distributed among the people, and it made a huge difference.
It’s a general problem with regression models on observational data. You have no idea what’s going on.
Madigan: “It’s the wild west out there.”
Wait, and it gets worse. It could be the case that within each group there males and females and if you partition by those you see that the more drugs they take the better again. Since a given person either is male or female, and either takes aspirin or doesn’t, this kind of thing really matters.
This illustrates the fundamental problem in observational studies, which is sometimes called Simpson’s Paradox.
[Remark from someone in the class: if you think of the original line as a predictive model, it’s actually still the best model you can obtain knowing nothing more about the aspirin-taking habits or genders of the patients involved. The issue here is really that you’re trying to assign causality.]
The medical literature and observational studies
As we may not be surprised to hear, medical journals are full of observational studies. The results of these studies have a profound effect on medical practice, on what doctors prescribe, and on what regulators do.
For example, in this paper, entitled “Oral bisphosphonates and risk of cancer of oesophagus, stomach, and colorectum: case-control analysis within a UK primary care cohort,” Madigan report that we see the very same kind of confounding problem as in the above example with aspirin. The conclusion of the paper is that the risk of cancer increased with 10 or more prescriptions of oral bisphosphonates.
It was published on the front page of new york times, the study was done by a group with no apparent conflict of interest and the drugs are taken by millions of people. But the results were wrong.
There are thousands of examples of this, it’s a major problem and people don’t even get that it’s a problem.
Randomized clinical trials
One possible way to avoid this problem is randomized studies. The good news is that randomization works really well: because you’re flipping coins, all other factors that might be confounders (current or former smoker, say) are more or less removed, because I can guarantee that smokers will be fairly evenly distributed between the two groups if there are enough people in the study.
The truly brilliant thing about randomization is that randomization matches well on the possible confounders you thought of, but will also give you balance on the 50 million things you didn’t think of.
So, although you can algorithmically find a better split for the ones you thought of, that quite possible wouldn’t do as well on the other things. That’s why we really do it randomly, because it does quite well on things you think of and things you don’t.
But there’s bad news for randomized clinical trials as well. First off, it’s only ethically feasible if there’s something called clinical equipoise, which means the medical community really doesn’t know which treatment is better. If you know have reason to think treating someone with a drug will be better for them than giving them nothing, you can’t randomly not give people the drug.
The other problem is that they are expensive and cumbersome. It takes a long time and lots of people to make a randomized clinical trial work.
In spite of the problems, randomized clinical trials are the gold standard for elucidating cause-and-effect relationships.
Rubin causal model
The Rubin causal model is a mathematical framework for understanding what information we know and don’t know in observational studies.
It’s meant to investigate the confusion when someone says something like “I got lung cancer because I smoked”. Is that true? If so, you’d have to be able to support the statement, “If I hadn’t smoked I wouldn’t have gotten lung cancer,” but nobody knows that for sure.
Define:
to be the treatment applied to unit
(0 = control, 1= treatment),
to be the response for unit
if
,
to be the response for unit
if
.
Then the unit level causal effect is , but we only see one of
and
Example: is 1 if I smoked, 0 if I didn’t (I am the unit).
is 1 or 0 if I got cancer and I smoked, and
is 1 or 0 depending on whether I got cancer while not smoking. The overall causal effect on me is the difference
This is equal to 1 if I got really got cancer because I smoked, it’s 0 if I got cancer (or didn’t) independent of smoking, and it’s -1 if I avoided cancer by smoking. But I’ll never know my actual value since I only know one term out of the two.
Of course, on a population level we do know how to infer that there are quite a few “1”‘s among the population, but we will never be able to assign a given individual that number.
This is sometimes called the fundamental problem of causal inference.
Confounding and Causality
Let’s say we have a population of 100 people that takes some drug, and we screen them for cancer. Say 30 out of them get cancer, which gives them a cancer rate of 0.30. We want to ask the question, did the drug cause the cancer?
To answer that, we’d have to know what would’ve happened if they hadn’t taken the drug. Let’s play God and stipulate that, had they not taken the drug, we would have seen 20 get cancer, so a rate of 0.20. We typically say the causal effect is the ration of these two numbers (i.e. the increased risk of cancer), so 1.5.
But we don’t have God’s knowledge, so instead we choose another population to compare this one to, and we see whether they get cancer or not, whilst not taking the drug. Say they have a natural cancer rate of 0.10. Then we would conclude, using them as a proxy, that the increased cancer rate is the ratio 0.30 to 0.10, so 3. This is of course wrong, but the problem is that the two populations have some underlying differences that we don’t account for.
If these were the “same people”, down to the chemical makeup of each other molecules, this “by proxy” calculation would work of course.
The field of epidemiology attempts to adjust for potential confounders. The bad news is that it doesn’t work very well. One reason is that they heavily rely on stratification, which means partitioning the cases into subcases and looking at those. But there’s a problem here too.
Stratification can introduce confounding.
The following picture illustrates how stratification could make the underlying estimates of the causal effects go from good to bad:
In the top box, the values of b and c are equal, so our causal effect estimate is correct. However, when you break it down by male and female, you get worse estimates of causal effects.
The point is, stratification doesn’t just solve problems. There are no guarantees your estimates will be better if you stratify and all bets are off.
What do people do about confounding things in practice?
In spite of the above, experts in this field essentially use stratification as a major method to working through studies. They deal with confounding variables by essentially stratifying with respect to them. So if taking aspirin is believed to be a potential confounding factor, they stratify with respect to it.
For example, with this study, which studied the risk of venous thromboembolism from the use of certain kinds of oral contraceptives, the researchers chose certain confounders to worry about and concluded the following:
After adjustment for length of use, users of oral contraceptives were at least twice the risk of clotting compared with user of other kinds of oral contraceptives.
This report was featured on ABC, and it was a big hoo-ha.
Madigan asks: wouldn’t you worry about confounding issues like aspirin or something? How do you choose which confounders to worry about? Wouldn’t you worry that the physicians who are prescribing them are different in how they prescribe? For example, might they give the newer one to people at higher risk of clotting?
Another study came out about this same question and came to a different conclusion, using different confounders. They adjusted for a history of clots, which makes sense when you think about it.
This is an illustration of how you sometimes forget to adjust for things, and the outputs can then be misleading.
What’s really going on here though is that it’s totally ad hoc, hit or miss methodology.
Another example is a study on oral bisphosphonates, where they adjusted for smoking, alcohol, and BMI. But why did they choose those variables?
There are hundreds of examples where two teams made radically different choices on parallel studies. We tested this by giving a bunch of epidemiologists the job to design 5 studies at a high level. There was zero consistency. And an addition problem is that luminaries of the field hear this and say: yeah yeah yeah but I would know the right way to do it.
Is there a better way?
Madigan and his co-authors examined 50 studies, each of which corresponds to a drug and outcome pair, e.g. antibiotics with GI bleeding.
They ran about 5,000 analyses for every pair. Namely, they ran every epistudy imaginable on, and they did this all on 9 different databases.
For example, they looked at ACE inhibitors (the drug) and swelling of the heart (outcome). They ran the same analysis on the 9 different standard databases, the smallest of which has records of 4,000,000 patients, and the largest of which has records of 80,000,000 patients.
In this one case, for one database the drug triples the risk of heart swelling, but for another database it seems to have a 6-fold increase of risk. That’s one of the best examples, though, because at least it’s always bad news – it’s consistent.
On the other hand, for 20 of the 50 pairs, you can go from statistically significant in one direction (bad or good) to the other direction depending on the database you pick. In other words, you can get whatever you want. Here’s a picture, where the heart swelling example is at the top:
Note: the choice of database is never discussed in any of these published epidemiology papers.
Next they did an even more extensive test, where they essentially tried everything. In other words, every time there was a decision to be made, they did it both ways. The kinds of decisions they tweaker were of the following types: which database you tested on, the confounders you accounted for, the window of time you care about examining (spoze they have a heart attack a week after taking the drug, is it counted? 6 months?)
What they saw was that almost all the studies can get either side depending on the choices.
Final example, back to oral bisphosphonates. A certain study concluded that it causes esophageal cancer, but two weeks later JAMA published a paper on same issue which concluded it is not associated to elevated risk of esophageal cancer. And they were even using the same database. This is not so surprising now for us.
OMOP Research Experiment
Here’s the thing. Billions upon billions of dollars are spent doing these studies. We should really know if they work. People’s lives depend on it.
Madigan told us about his “OMOP 2010.2011 Research Experiment”
They took 10 large medical databases, consisting of a mixture of claims from insurance companies and EHR (electronic health records), covering records of 200 million people in all. This is big data unless you talk to an astronomer.
They mapped the data to a common data model and then they implemented every method used in observational studies in healthcare. Altogether they covered 14 commonly used epidemiology designs adapted for longitudinal data. They automated everything in sight. Moreover, there were about 5000 different “settings” on the 14 methods.
The idea was to see how well the current methods do on predicting things we actually already know.
To locate things they know, they took 10 old drug classes: ACE inhibitors, beta blockers, warfarin, etc., and 10 outcomes of interest: renal failure, hospitalization, bleeding, etc.
For some of these the results are known. So for example, warfarin is a blood thinner and definitely causes bleeding. There were 9 such known bad effects.
There were also 44 known “negative” cases, where we are super confident there’s just no harm in taking these drugs, at least for these outcomes.
The basic experiment was this: run 5000 commonly used epidemiological analyses using all 10 databases. How well do they do at discriminating between reds and blues?
This is kind of like a spam filter test. We have training emails that are known spam, and you want to know how well the model does at detecting spam when it comes through.
Each of the models output the same thing: a relative risk (causal effect estimate) and an error.
This was an attempt to empirically evaluate how well does epidemiology work, kind of the quantitative version of John Ioannidis’s work. we did the quantitative thing to show he’s right.
Why hasn’t this been done before? There’s conflict of interest for epidemiology – why would they want to prove their methods don’t work? Also, it’s expensive, it cost $25 million dollars (of course that pales in comparison to the money being put into these studies). They bought all the data, made the methods work automatically, and did a bunch of calculations in the Amazon cloud. The code is open source.
In the second version, we zeroed in on 4 particular outcomes. Here’s the $25,000,000 ROC curve:
To understand this graph, we need to define a threshold, which we can start with at 2. This means that if the relative risk is estimated to be above 2, we call it a “bad effect”, otherwise call it a “good effect.” The choice of threshold will of course matter.
If it’s high, say 10, then you’ll never see a 10, so everything will be considered a good effect. Moreover these are old drugs and it wouldn’t be on the market. This means your sensitivity will be low, and you won’t find any real problem. That’s bad! You should find, for example, that warfarin causes bleeding.
There’s of course good news too, with low sensitivity, namely a zero false-positive rate.
What if you set the threshold really low, at -10? Then everything’s bad, and you have a 100% sensitivity but very high false positive rate.
As you vary the threshold from very low to very high, you sweep out a curve in terms of sensitivity and false-positive rate, and that’s the curve we see above. There is a threshold (say 1.8) for which your false positive rate is 30% and your sensitivity is 50%.
This graph is seriously problematic if you’re the FDA. A 30% false-positive rate is out of control. This curve isn’t good.
The overall “goodness” of such a curve is usually measured as the area under the curve: you want it to be one, and if your curve lies on diagonal the area is 0.5. This is tantamount to guessing randomly. So if your area under the curve is less than 0.5, it means your model is perverse.
The area above is 0.64. Moreover, of the 5000 analysis we ran, this is the single best analysis.
But note: this is the best if I can only use the same method for everything. In that case this is as good as it gets, and it’s not that much better than guessing.
But no epidemiology would do that!
So what they did next was to specialize the analysis to the database and the outcome. And they got better results: for the medicare database, and for acute kidney injury, their optimal model gives them an AUC of 0.92. They can achieve 80% sensitivity with a 10% false positive rate.
They did this using a cross-validation method. Different databases have different methods attached to them. One winning method is called “OS”, which compares within a given patient’s history (so compares times when patient was on drugs versus when they weren’t). This is not widely used now.
The epidemiologists in general don’t believe the results of this study.
If you go to http://elmo/omop.org, you can see the AUM for a given database and a given method.
Note the data we used was up to mid-2010. To update this you’d have to get latest version of database, and rerun the analysis. Things might have changed.
Moreover, an outcome for which nobody has any idea on what drugs cause what outcomes you’re in trouble. This only applies to when we have things to train on where we know the outcome pretty well.
Parting remarks
Keep in mind confidence intervals only account for sampling variability. They don’t capture bias at all. If there’s bias, the confidence interval or p-value can be meaningless.
What about models that epidemiologists don’t use? We have developed new methods as well (SCCS). we continue to do that, but it’s a hard problem.
Challenge for the students: we ran 5000 different analyses. Is there a good way of combining them to do better? weighted average? voting methods across different strategies?
Note the stuff is publicly available and might make a great Ph.D. thesis.







hey Cathy, you are doing an awesome job with blogging about the class!
LikeLike
I met David Madigan once briefly, and he seemed very cool.
LikeLike